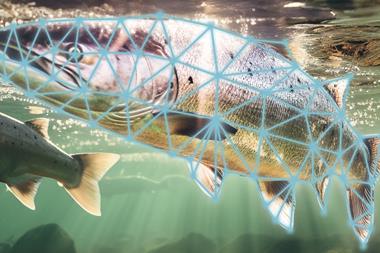
The genes of the monkeyface prickleback could hold the secret to more sustainable aquaculture.
Commonly referred to as the monkeyface eel, the monkeyface prickleback (Cebidichthys violaceus) lives in the rocky waters off the US west coast and eats a vegetarian diet of red and green algae. Preferring to lurk in rocky reefs and tide pools, it’s an unusual-looking species with two small pectoral fins hanging like floppy ears near its head and a dorsal fin that winds down its back.
Herbivorous fish only make up 5% of total fish species on the planet, but when it comes to aquaculture, they are extremely significant – they are more sustainable and less expensive to farm than carnivorous species.
This caught the attention of Donovan German, Associate Professor of Ecology and Evolutionary Biology at the University of California, Irvine, researcher Joseph Heras and their colleagues. Curious to discover how the monkeyface prickleback survives on food containing a low level of lipids, they sequenced and assembled a high-quality genome for the fish and obtained a blueprint for what’s required to be a herbivore.
Joseph Heras extracted genomic DNA from the fish’s muscle tissue and used two technologies to sequence the genome. A specific bioinformatic software called “quickmerge,” developed by study co-authors Mahul Chakraborty and J.J.Emerson, was then used to put the genome together, while the transcriptome – genes expressed from a given cell or tissue – was sequenced from nine different tissues (brain, gill, testes, heart, liver, pyloric caeca, proximal intestine, middle intestine and spleen). These helped the team annotate the genome and see how the different tissues operate differently from one another, Joseph Heras said.
“We identified three copies of amylase, an enzyme which breaks down starch,” Donovan German commented.
“We also compared the amylase genes from the monkeyface prickleback with other prickleback fishes. We found that one of the amylase genes in the monkeyface prickleback, which we call amy2b, is very different from those in other prickleback fishes. We also looked at lipases, an enzyme which breaks down lipids, and found four carboxyl ester lipase genes in a row on the same chromosome. This is interesting because the algae that the monkeyface prickleback consumes contains a low amount of lipids overall. The finding suggests that the species has invested in obtaining that lipid, even if it’s scarce in its diet. In other words, it has adapted to be very efficient at breaking down lipids, even though lipids comprise just five percent of the algae’s composition.”
Digestive specialisation
According to the team, this is a compelling example of what they call ‘digestive specialisation’ in the genome. The discovery is also promising because it highlights the genomic blueprints of what is required for fish to digest plant material, and could lead to a new source of protein for human consumption. It could also be a particularly suitable target for aquaculture. One major issue for the industry is what to feed the fish being raised. Being able to give them vegetation will save money and be better for the environment.
“Most of the fish we culture for food, such as salmon and sea bass, are carnivorous and do poorly when fed plant material, particularly plant-based lipids. That’s why they are reared on fishmeal-based diets but this isn’t sustainable in the long-term,” Joseph Heras explained.
“The monkeyface prickleback provides a way towards developing sustainable, plant-based feeds on which we can grow fishes in culture, either by finding other “tasty” fish with similar capabilities as the monkeyface prickleback, or by inserting monkeyface prickleback genes into the fish we want to eat, unlocking new abilities in them to digest and grow on plant-based feeds.”
“The monkeyface prickleback is also a delicacy on the central California coast, and there is interest in culturing this species for human consumption. Having the genome better prepares for that,” Donovan German added.
Fish feed has become increasingly limited and costly in the midst of climate change, while an increasing global population is raising the demand for protein sources, especially sustainable resources, says German. But having a high quality, herbivorous fish genome means that German, Heras and their colleagues now know what to look for in other fish genomes if aquaculture wants to culture them for food.
Reduces pollution and costs
“The monkeyface prickleback is adapted to eating algae found within tide pools and shallow rocky areas,” Joseph Heras said.
“This gives us a good example of what genes, gene expression patterns, or gene copies would be necessary to make a living off plant material, the lipids in particular. Lastly, using plant-based food ingredients reduces pollution and costs less.”
He said that the likelihood of the monkeyface prickleback having a future in aquaculture on a massive scale is hard to tell because the species thrives in cold waters and grows slowly. But, he says, if these conditions can be met, and don’t require much energy and costs, the species could be farmed in areas such as the central California coast. So far, the team has had enquiries from various institutions on the lipid digestion and metabolism of herbivorous fish, and is now assembling the genomes of another herbivorous prickleback species (Xiphister mucosus) and two omnivorous pricklebacks (Xiphister atropurpureus and Phytichthys chirus).
This is expected to enable the team to make stronger inferences of what is required for herbivory or omnivory and better understand the nutritional requirements of fishes. It may also be used to improve the farming of species that have these types of diet specialisations, and manage the habitats along the California coast where these species live. Work is also underway to delve further into the biochemistry of the lipase proteins in the monkeyface prickleback, to learn more about what it takes to digest plant-based lipids.
The team also believes that its work will lead to the discovery of more potential aquaculture species. One of its next steps is to apply the information and knowledge it’s gained so far towards new projects with other species.
“As the price of genomic sequencing decreases, this provides the opportunity to sequence more genomes of species of interest, especially herbivorous fishes, and compare these genomes to the monkeyface prickleback genome. Thus, we can find tasty fish with the capability to thrive on plant-based feeds,” Joseph Heras said.
“Also, with the improvement in computational resources, I can definitely see more comparative genomic methods used to show the genetic requirements of herbivory in fishes. I’m fortunate to have landed an assistant professor (tenure track) position at California State University, San Bernardino,” he continued.
“I just started this fall 2020, so Donovan and I will continue to collaborate on these prickleback genomic and transcriptomic projects.”